The Urobiome and Its Role in Overactive Bladder
Article information
Abstract
Urine is no longer considered to be sterile. After the existence of the microbiome was revealed through metagenomic analysis using next-generation sequencing, the relationship between characteristics of the microbiome and diseases have been studied and published in various journals. A microbiome exists in the urinary tract and is associated with urinary tract infection, malignancy of the genitourinary tract, and lower urinary tract symptoms. Based on the urine sampling method, sampling site, culture method, and sex, the characteristics of the microbiome vary. Most of the Lactobacillus species are identified mainly in women, and various other species are identified in men. These microorganisms can cause or prevent various diseases. Variations in the microbiome are seen in those with and without disease, and an asymptomatic status does not indicate the absence of microbes. This microbiome has been implicated in a variety of lower urinary tract symptoms and diseases, in particular, overactive bladder. The microbiome differs between patients with urgency and urge urinary incontinence and healthy individuals. There are many aspects of the microbiome yet to be studied in relation to other lower urinary tract symptoms.
INTRODUCTION
The term microbiota was first described by Lederberg and McCray, and ‘microbiome’ refers to the entire habitat, including the microorganisms such as bacteria, archaea, lower and higher eukaryotes, viruses, and their genome and surrounding environment [1]. Through next-generation sequencing (NGS) [2], in addition to the pathological strains previously confirmed through culture tests, many normal flora (microbiota) were identified. These microbial environments keep humans healthy, prevent diseases, and play a role in homeostasis. Changes in these environments cause diseases by disrupting homeostasis.
Microbiomes are commonly found in the skin, placenta, mammary glands, seminal fluids, uterus, ovarian follicles, lung, saliva, oral mucosa, conjunctiva, biliary tract, and gastrointestinal tract system [3,4]. In 2015, Kogan et al. [5] announced that human urine was not sterile. Many researchers have confirmed the existence of microbiota in the human urinary tract through metagenomics (DNA-dependent) and metaculturomics (culture-dependent) technologies. Previously, it was believed that urine is sterile and that uropathogens invaded and caused urinary tract infections (UTIs). However, as the existence of the microbiome in urine-the urobiome-was confirmed, a new paradigm for the pathogenesis of diseases was suggested. The urobiome plays a role in preventing infections caused by pathogens, forms a barrier, and is necessary to maintain a healthy urinary tract; it has helped us gain a deeper understanding of various diseases. Currently, the microbiome is being studied in various fields of urology, such as UTI, interstitial cystitis, and genitourinary tract malignancy.
In addition to UTIs, the notion that the microbiome is associated with various urinary tract diseases is gaining popularity, and its relationship with various lower urinary tract symptoms is being studied [6,7]. A precise understanding of the urobiome is necessary to understand and treat urinary diseases in relation to the microbiome.
Among the disease groups with lower urinary tract symptoms, there are diseases whose prevalence is increasing owing to aging of the population, and the representative disease among them is overactive bladder (OAB). In the absence of an obvious infection, overactive bladder syndrome is characterized by urinary urgency, with or without urgency urinary incontinence, and is usually accompanied by frequency and nocturia [8]. The cause of OAB remains unknown. However, obesity, caffeine intake, constipation, diabetes, poor mobility, and chronic pelvic pain may be risk factors [9,10]. The diagnosis is mainly based on the signs and symptoms after UTI or neurological deficit has been excluded. It has a prevalence of 9%–43% in women and 7%–27% in men, and the prevalence tends to increase with age. Originally, detrusor overactivity (DO) was believed to be the major pathophysiological cause of OAB. DO can either be ‘myogenic,’ where the detrusor muscle contracts due to autonomic stimulation, or ‘neurogenic,’ where detrusor contraction is induced by the urgency signal released by the central nervous system. Recently, as the importance of the urothelium and suburothelium has been emphasized; different approaches and analyses for pathophysiology have been developed. In terms of the urothelium/suburothelium, various elements are being studied, including the urinary microbiota or urobiome. Peyronnet et al. [11] reported the pathophysiological factors affecting OAB are metabolic syndrome, affective disorder, sex hormone deficiency, urinary microbiota, gastrointestinal functional disorder, and subclinical autonomic nervous system dysfunction. In this regard, we aimed to investigate the urobiome and its relevance in OAB.
WHAT IS UROBIOME?
The urobiome refers to the microbiome of the urinary tract [12]. Since there was a strong assumption that urine is sterile in healthy people, the microbiome of urine was not included in the initial Human Genome Project. However, as the microbiomes of other organs have been attracting attention, and various microbiomes have been identified in various environments since then, there is an increase in the research regarding the microbiome of the urinary tract and its involvement in urologic diseases. However, studies show different results depending on the study conditions, and there are some difficulties in matching these conditions for comparisons between studies. There are several criteria to be considered when conducting research, which are: (1) collection method, (2) sex, (3) age, and (4) specific diseases of each organ of the urinary tract.
METHODS USED FOR URINARY MICROBIOME ANALYSIS
Microorganism detection methods can be divided into culture-based and sequencing-based methods. Traditionally, microorganisms have been identified by a standard urine culture method using MacConkey agar or blood agar. Rapid-growing and aerobic microorganisms were mainly identified. However, there is a disadvantage to culture-based methods as anaerobic and slow-growing microorganisms cannot be cultured.
The existence of the microbiome was confirmed by metagenomic analysis using a method called NGS [13]. It is a sequencing-based method that identifies bacteria through DNA sequences rather than the traditional method of culturing bacteria. It is divided into 2 types, amplicon sequencing using ‘marker genes’ such as 16S rRNA subunit with 9 hypervariable regions, and shotgun sequencing to analyze the entire microbiome sequence [14]. The disadvantage of the sequencing methods is that the viability of the microbes cannot be confirmed. An enhanced quantitative urine culture (EQUC), as a traditional culture-based method, can identify bacteria that were previously difficult to identify and compensate for such shortcomings [15]. An EQUC is performed by incubating the sample that was cultured using blood, chocolate, and colistin-nalidixic acid agar for 48 hours at 35°C A disadvantage of EQUC is that it identifies a broad range of bacteria, requiring interpretation of the results between patients and healthy people. Therefore, it is recommended that 16S rRNA gene sequencing and EQUC be performed simultaneously.
PROBLEMS WITH URINE COLLECTION
The existence of a microbiome in urine has been established by NGS and has been drawing attention from researchers. The detection of the diversity of the microbiome is impacted by various factors, one often mentioned is the sample collection method [16]. Since the microbiome is greatly influenced by the surrounding environment, different results are achieved depending on where the sample used for the study was collected. Pohl et al. [17] reported that a different microbiome was identified in the urine that passed through both the bladder and urethra through self-voiding compared with the sample that bypassed the urethra through the catheter. Wolfe and Brubaker [18], ‘Is it self voiding?’ ‘Is it midstream?’ ‘Is it Catheterization?’ ‘Is it a blade puncture?’ Different results were obtained depending on the differences in the urinary tract through which the sample urine passed [18] (Fig. 1). In another study, there was no difference in the microbiome results between the first and mid streams [19]. This means that the microbiome in the urethral environment and the microbiome of the bladder are different, and whether the microbiome of the urethra is included impacts the results. Regarding the upper ureter, its microbiome is affected by changes in the microbial environment at the kidney level. Studies have shown that a decrease in estimated glomerular filtration rate reduces microbiome diversity [20].
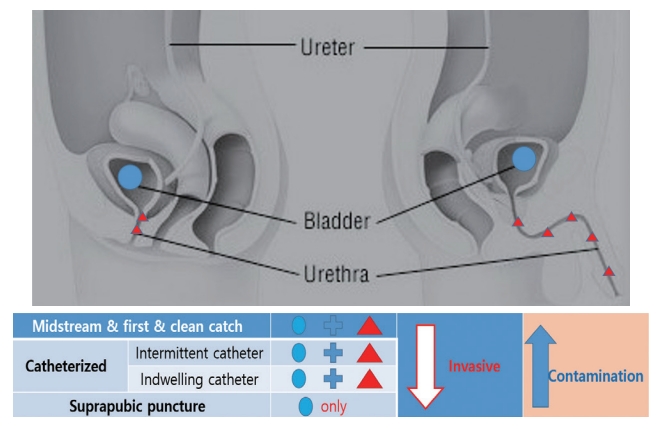
Differences in urine collection methods in relation to the urobiome (invasiveness and contamination). In the case of suprapubic puncture, only the microbiome of the bladder was identified, and the invasiveness was the highest. Conversely, self-voiding methods (first catch urine, midstream urine, and clean catch urine) are less invasive but are contaminated with microbiota from the urethral flora.
Although the microbiome differs from individual to individual, depending on odor, method of sample collection, and sex, there is one important thing in common: there is a difference between asymptomatic and symptomatic patients. It is important to remember that an asymptomatic status does not indicate the absence of disease.
DIFFERENCES IN THE URINARY MICROBIOME BETWEEN MEN AND WOMEN
The male and female reproductive microbiomes are different [21]. In a recent study, it was confirmed that the microbiome found in the genital area of men and women was different before sexual intercourse, but was similar after sexual intercourse [22]. The urine microbiome also differs between men and women [17,23]. There are several studies that analyzed urine samples collected from healthy men and women using various methods, such as clean catch urine, transurethral catheter urine, suprapubic puncture, midstream urine, and first catch urine; the common discovery was that Lactobacillusmethods. However, in men, a heterogeneous result was obtained based on the sampling method (Table 1).
AGE
There is a controversy about age as a factor influencing the microbiome, there are reports of a lack of relevance [24] and correlation [3,25,26]. When checked if microbiome is directly affected by age, it was found that there was no significant age-related change; however, there was a difference between the young and old population [27,28]. Rather than considering age as a factor, the change in microbiome can be considered to be a secondary change associated with aging. Various changes in the body as a result of aging can cause changes in the environmental aspect of the microbiota, which affects microbiome diversity and commensal microorganisms, which can be seen not only in people with diseases but also in healthy people. In women, hormonal changes occur because of aging; therefore, menopause is an important criterion for change in the microbiome. It is evident that environmental changes in the vaginal microbiome caused by female hormones cause changes in the urobiome. According to several studies, Lactobacillus is abundant in the urine of premenopausal women, and less so in postmenopausal women. In addition, Mobiluncus spp. were found abundantly in postmenopausal women [24].
DISEASE-RELATED CHANGES IN THE URINARY MICROBIOME IN WOMEN IN RELATION TO CHANGES IN THE GUT MICROBIOME
Some urologic diseases in women are a result of the imbalance in the urobiome, such as: OAB syndrome, urge urinary incontinence (UUI), interstitial cystitis/bladder pain syndrome, asymptomatic bacteriuria, and neurogenic bladder dysfunction. Although there is no clear mechanism or association yet, many studies have shown a link between the disease and the microbiome [14,25,29-33]. Lactobacillus, which was identified as an important contributor in these diseases, changes the vaginal flora while also changing the gut microbiome through oral probiotics and causes diseases such as diarrhea in the gastrointestinal tract. These changes play a role in the prevention of UTIs. A recent study also reported that changes in the gut microbiota affect OAB and daily urgency [34]. There was a difference in the gut microbiota between the group with OAB symptoms and urgency and the asymptomatic group. It is common to consider the urinary microbiome to be an ascending infection of the gut. The urinary microbiome was found to be 62.5% similar to the gut microbiome and 32% similar to the vaginal microbiome; therefore, they are closely related to each other [16,35-37].
URINARY BIOMARKERS AND MICROBIOME IN OVERACTIVE BLADDER
In OAB syndrome, various urine biomarkers were extensively studied to distinguish phenotypes along with the diagnosis [3,4,38,39]. A representative example is nerve growth factor (NGF) and brain-derived neurotrophic factor, a neurotrophin that is produced in the urothelium or smooth muscle cells is widely used as a marker of lower urinary tract dysfunction. According to Suh et al. [40], urinary NGF increases in patients with OAB, hence is considered an important biomarker for the treatment of OAB. Another biomarker is microRNA in genomics. miRNAs have also been used as biomarkers for urothelial carcinoma in the urological field [41,42]. Some microRNAs, such as miR-103a-3p, miR-10a-5p, and miR-199a-3p, have been used as urinary biomarkers of DO or underactivity [43].
The interdependence between antimicrobial peptides (AMPs), specifically beta-adhesin AMP, and the microbiota increased the risk of UTI after surgery [44]. In addition, levels of prostaglandin E2 (PGE2), an inflammatory marker also increases in OAB [45]. PGE2 is also associated with bladder capacity, and its levels are significantly decreased after administration of intravesicular botulinum toxin A injection [46]. Furthermore, high levels of PEG2 are observed in UUI and is considered as a potential target for future treatment. Urinary adenosine triphosphate (ATP) is also involved in contraction and relaxation of the detrusor muscle, and in patients with OAB, it may be overexpressed or its concentration in urine may change [47,48]. Urinary b3-adrenoceptor (B3-AR) is involved in detrusor relaxation, and the Trp64Arg polymorphism in B3-AR is closely related to DO [49]. The urinary microbiome shows characteristics specific to the disease, a large number of specific strains are identified in diseases such as UUI across various samples. As a result, it can be used as a useful biomarker of urine if it is further standardized and researched based on a large number of subjects (Table 2).
CORRELATION BETWEEN OVERACTIVE BLADDER SYMPTOMS AND MICROBIOME
In the absence of a UTI, the signs and symptoms (urgency complaints) are diagnosed as a representative bladder storage disorder. However, as the urothelium/suburothelium is known to play an important role in pathophysiology of OAB, studies about the correlation between the symptoms and microbiome of OAB are being conducted. According to Peyronnet et al. [11], OAB phenotypes include metabolic syndrome, affective disorder, sex hormone deficiency, functional gastrointestinal disorder, autonomic nervous system dysfunction, and ‘urinary microbiota’ depending on the pathophysiological factors . It is also predicted that brain-bladder-microbiota amblyopia exists as the gut microbiota forms the brain-gut-microbiota axis.
However, it has not yet shown any significant results for each symptom, and the relationship with the microbiome has been revealed in some symptoms. In general, symptoms of OAB include frequency, urgency, nocturia, and urge incontinence. The role of the microbiome in each can be summarized as shown in Table 2. Studies have shown that there is a difference in the microbiome of patients diagnosed with OAB, regardless of the symptoms. According to Curtiss et al. [50], there is a study showing that Proteus was significantly abundant in patients with OAB compared to healthy people, and at the same time, Lactobacillus was less prevalent (Table 3).

Microbial strains identified in urine samples collected from individuals with overactive bladder symptoms
Frequency
There are negligible number of papers that mention the correlation between frequency and the microbiome. However, when examining each symptom cluster using EQUC in patients UTI, the presence of Lactobacillus or Klebsiella among the symptom clusters with frequency symptoms were significantly prevalent compared to the asymptomatic group [51]. We concluded that more clear research is needed in this area.
Nocturia
Although the correlation between nocturia symptoms and the microbiome is unknown, Holland et al. [52] reported that nocturia was related to the International Prostate Symptom Score (IPSS). Lachnospiraceae Blautia in urinary operational taxonomy units has bothersome symptoms, is associated with severity, has protective effects in IPSS, and is also associated with nocturia.
Urgency
A study compared urgency with Overactive Bladder Symptom Score using the gut microbiome and not urine microbiome. In the OAB group, bacterial diversity was low, the Bifidobacterium group was low, and the Faecali bacterium group was abundant.
Urinary incontinence
Among the symptoms associated with OAB, urinary incontinence is the most studied and is the best-known symptom of microbiome imbalance [53]. Although incontinence and, in particular, UUI, are not essential symptoms of OAB, various studies have considered their relationship with DO [25,29,30,33,34,54-59].
Pearce et al. [29] compared patients with and without UUI symptoms and confirmed a difference in prevalence of Gardenerella and Lactobacillus. In the UUI group, presence of Gardnerella was high and Lactobacillus was low. In the same study, Lactobacillus spp. levels varied in the control group, Lactobacillus crispatus was high while Lactobacillus gasseri was low. In UUI, vaginal estrogen therapy in postmenopausal women resulted in a decrease in microbial diversity, a change in Lactobacillus levels in urine, and a change in UUI [60].
In addition to these strains, E. coli and some Gardnella vaginalis strains, classified as uropathogenic microorganisms, induce the release of calcium ions or ATP from the urothelial epithelium and are involved in the contraction of bladder smooth muscle cells, causing incontinence. In contrast, Lactobacillus has been reported to inhibit this process [61]. Furthermore, in patients with refractory UUI with recurrent UTI, diverse microbiota exists, but if colonization continues, the disease persists, so appropriate intervention is necessary [62].
There are studies on the role of the urobiome not only with respect to the presence or absence of UUI symptoms, but also to symptom severity. Karstens et al. [56] cultured bladder microbiomes of healthy individuals and individuals with UUI. The study identified 14 strains and established that there are differences in the relative abundance in healthy people and patients with UUI, and that the microbiome in individuals with UUI differed according to severity.
LIMITATION OF OAB RESEARCH USING THE MICROBIOME
There are several problems in studying the association between the symptoms of OAB, pathophysiology, and microbiome:
(1) It can be concluded that there is some association, but in some cases, there is uncertainty regarding the predominance of a specific microbiome in the diseased and nondiseased individuals, for example: Lactobacillus predominance does not differ between adult women with mixed urinary incontinence and age-matched asymptomatic women, but some members of the genus Lactobacillus might be associated with urinary symptoms [28].
(2) In many cases, the collected data are not suitable for studying the urobiome and data on a specific microbiome are insufficient or absent. Public databases are inadequate for studies of the urobiome and its relationship with bladder health and diseases because these databases lack urobiome-specific genomes [63]. This is due to the fact that the dominant microbiome is different under various environmental conditions. However, since there is a lack of standardization, several factors, such as the method and timing of sample collection, sex, and comorbidities, must be considered. Only when the research is conducted by considering all these aspects will it be possible to derive more reliable results on the relevance of a specific microbiome with respect to diseases or symptoms.
(3) It is necessary to standardize the sample, especially in men, and the urethral environment must be considered. Men with more severe urinary symptoms are more likely to have detectable bladder bacteria than those with few severe or no symptoms. Voided urine does not adequately characterize the male bladder urobiome and catheterized urine should be used instead [64].
(4) Studies with large sample sizes are scarce. More extensive and large-scale data is needed on samples of healthy and diseased individuals.
CONCLUSIONS
The discovery of the microbiome using NGS has completely changed the disease paradigm. Urine is not considered sterile anymore. In situations where association between various diseases of the urinary tract and the role of the urothelium are emerging, various clinical aspects of OAB have been shown closely related to the urobiome. Various approaches and considerations, such as the gut microbiome and vaginal flora, are required to study the relationship between the urobiome and OAB. However, since the size of the study and the sample size was not large enough, a clearer and more accurate analysis is needed through a larger-scale and harmonized research method.
Notes
Fund/Grant Support
This paper was written as part of Konkuk University’s research support program for its faculty on sabbatical leave in 2019.
Conflict of Interest
No potential conflict of interest relevant to this article was reported.
AUTHOR CONTRIBUTION STATEMENT
·Conceptualization: CH
·Data curation: BSR
·Formal analysis: BSR, CH
·Funding acquisition: BSR, CH
·Methodology: CH
·Project administration: CH
·Visualization: BSR
·Writing - original draft: BSR
·Writing - review & editing: BSR, CH


