Human Endogenous Retroviruses: Friends and Foes in Urology Clinics
Article information
Abstract
Human endogenous retroviruses (HERVs) are originated from ancient exogenous retroviruses, which infected human germ line cells millions of years ago. HERVs have generally lost their replication and retrotransposition abilities, but adopted physiological roles in human biology. Though mostly inactive, HERVs can be reactivated by internal and external factors such as inflammations and environmental conditions. Their aberrant expression can participate in various human malignancies with complex etiology. This review describes the features and functions of HERVs in urological subjects, such as urological cancers and human reproduction. It provides the current knowledge of the HERVs and useful insights helping practice in urology clinics.
INTRODUCTION
Human endogenous retroviruses (HERVs) have infected and integrated into the human germline, mostly after the divergence of the New and the Old World monkeys [1-4]. Over millions of years, the human genome has accumulated hundreds of thousands of HERVs and related retrotransposons, comprising approximately 8% of the human genome [5,6].
The genomic structure of HERV is relatively simple, containing 4 primary viral genes of gag, pro, polymerase (pol), and envelop (env), flanked by 2 long terminal repeats (LTRs) [7]. The gag encodes the viral matrix, capsid, and nucleocapsid proteins. The pro specifies protease and the pol contains coding sequences of reverse transcriptase (RT), ribonuclease H, and integrase. The ENV protein product of the env gene has a surface and a transmembrane subunit. The LTRs have regulatory functions for both viral and host genes as well as retrotranspositions. The HERV-K group presents 2 additional accessory proteins, Np9 and Rec, translated from alternatively spliced env transcripts [8,9]. Type I human mouse mammary tumor virus like-2 (HML-2) generates Np9, and both type II HML-2 and type II HML-10, Rec, using specific alternative splicing sites [10].
HERVs are largely repressed due to mutations accumulated throughout evolution and epigenetic modifications such as DNA modification. Nevertheless, some distinct HERVs are competent to produce transcripts or proteins, which participate in crucial processes such as transcription in the nucleus, immune system activation, and placenta development [11-13]. HERV sequences in the genome can serve as regulatory elements that affect gene expression by interacting with various transcription factors (TFs), including some key players in physiological and pathological processes [14]. Also, failure to repress HERVs or their dysregulated expression is associated with numerous human diseases and disorders, including various cancers and infertility, which involve many critical molecular, cellular, and physiological pathways [15-21]. The unwanted activation of HERVs can be induced by inflammation, endocrine imbalance, exogenous viral infections, chemical exposures, and other environmental conditions [4,22-24]. Understanding the way HERVs function for human biology and clinical status supports the development of efficient therapeutic tools for treating various human illnesses [25-30].
HERVs are classified into many groups, generally based on sequence similarity, predicted proviral features such as nearby LTRs, closely-related exogenous retroviruses, species in which they were first identified, and their usage of tRNA primers [31]. For example, HERV designates ERV initially seen in the human genome, and HERV-K indicates ERVs that use a lysine tRNA. The HERV-K group is further classified into multiple subgroups, such as HML-2, based on their taxonomic backgrounds [32].
In this study, we compile literatures, which describe specific features and functions of HERVs in urological subjects, such as urological cancers, and male and female reproduction (Fig. 1). The insights from emerging implications of HERVs in general urology and human reproduction help understand the roles of HERVs in human physiology and evolution.
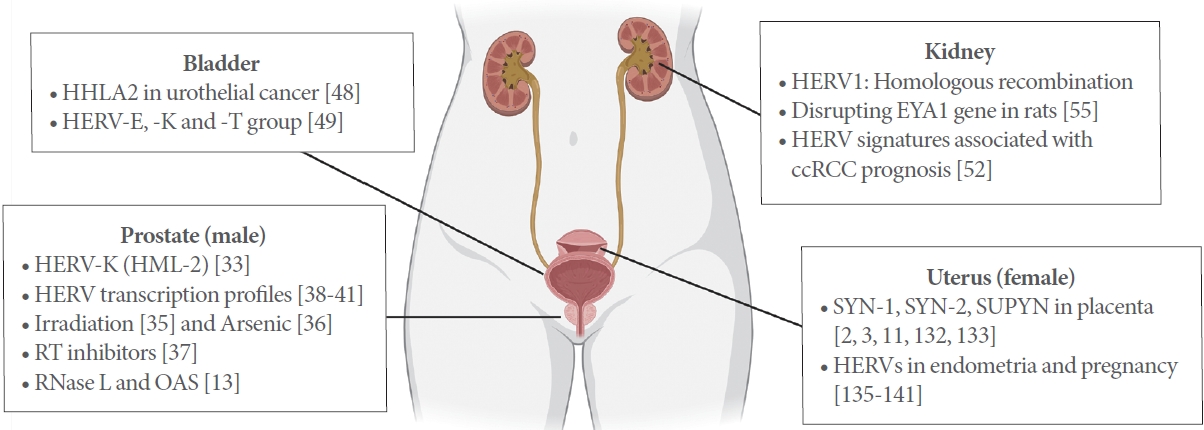
Human endogenous retroviruses (HERVs) in the human urological system. HHLA2, HERV-H long terminal repeat-associating protein 2; EYA1, eyes absent homolog 1; ccRCC, clear cell renal cell carcinoma; HML-2, human mouse mammary tumor virus like-2; RT, reverse transcriptase; OAS, 2’,5’-oligoadenylate synthetases; SYN-1, syncytin-1; SYN-2, syncytin-2; SUPYN, suppressyn. Adapted from BioRender.com with permission.
HERVs IN UROLOGICAL CANCERS
Prostate Cancer
HERVs are actively transcribed in various cancers, including prostate cancer [2,3]. HERV-K (HML-2) transcripts are detected in prostate cell lines such as LNCaP, DU145, PC3, and VCaP [33]. Splicing variants for env mRNAs are detected. Most transcript sequences matched with internal regions of HERV-Ks are short, but those from the LTRs are relatively intact. They arise from multiple HERV-K loci, and proviruses are also detected. Transcripts from antisense strands and both strands of solo LTRs are also detected. Multiple HERV-K loci are actively transcribed in prostate cancer cell lines, and either strand of viral genes can serve for transcription.
HERV expression in prostate cancer cells is affected by environmental factors such as irradiation and toxic materials. Ionizing radiation transiently increases HERV-K transcription in prostate cancer cell lines [34]. The transcript level peaks at 24 hours after a single dose of gamma-irradiation, ranged from 2.5 to 20 Gy, and returns to baselines within 72 hours. Irradiation triggers expressions of immunity-mediated genes, to which HERV expression may contribute [35]. CAsE-PE cells are an arsenic-transformed human prostate epithelial cell line containing oncogenic mutations in KRAS [36]. In these cells, the mutated KRAS is amplified, and KRAS viral fusion transcripts are detected, primarily mapped to LTR and HERVs. The KRAS viral fusion transcripts are heterogenous, suggesting multiple retroviral integrations. Therefore, environmental toxins such as arsenic may activate HERVs and possibly involve in carcinogenesis.
High RT activity is often reported in cancer cells and nonnucleoside RT inhibitors to treat HIV infection exert a robust cytostatic and differentiating activity in several cancers, presumably inhibiting HERV RT activity. The differentiating activity of RT inhibitors may help improve conventional treatments in hormone-refractory prostate cancers [37]. It is suggested that widely-applied treating drugs for HIV infection could be used as anticancer therapeutic tools.
RNase L exerts antiviral and antitumor activities by binding to the allosteric effectors 5´-phosphorylated, 2´, 5´-linked oligoadenylates (2-5A), which is produced by interferon (IFN)-inducible 2´, 5´-oligoadenylate synthetases (OAS). RNase L mutation is implicated as a risk factor for prostate cancer, and it is reported that the HERV env RNAs bind and activate OAS in prostate cancer cell line PC3 cells [13]. Therefore, high expression of HERV env RNAs in prostate cancers may contribute to the induction of antiviral and antitumor activities.
The prostate-specific antigen (PSA) is the primary diagnostic biomarker for prostate cancer in clinical use, but it lacks specificity and sensitivity [38]. HERV-driven RNAs have been shown as a potential alternative for prostate cancer diagnosis [39]. HERV-K env protein level is commonly increased in prostate tumors. HERV-K gag transcripts levels in peripheral blood mononuclear cells are higher in prostate cancer patients than in healthy subjects, and associated with IFN-γ [40]. High gag expression is associated with the diagnosis with prostate cancer, especially in older men and smokers, who tend to develop a more aggressive form of the disease. Combining noninvasive HERV-K testing with PSA testing may improve the efficacy of prostate cancer detection. High-density Affymetrix chip targeting 2,690 distinct proviruses and 2,883 solo LTRs of the HERVW, HERV-H, HERV-E 4.1, HERV-FRD, HERV-K HML-2 and HERV-K HML-5 families, unveil the expression of 1,718 HERV loci in a wide range of tissues [41]. The study identifies putative prostate cancer biomarkers and prostate cancer-specific HERV transcription regulation, such as proviral splicing events and methylation-dependent epigenetic regulation.
The LTRs of HERVs often regulate the expression of nearby genes. The expression of HML-2, the most biologically active subgroup of the HERV-K family, is associated with many cancer types, including prostate cancer. The LTRs of HML-2 have been classified into 3 subgroups: LTR5A, LTR5B, and LTR5Hs. It is reported that the LTR5Hs group is clearly separated from the other 2 groups in a phylogenetic tree and shows the most potent promoter activity, which relies on two p53 binding sites located between -263 and 0 [14]. The direct binding of the p53 protein is evidenced by chromatin immunoprecipitation (ChIP) and CUT&Tag experiments. p53 is a tumor suppressor protein, which regulates the expression of genes related to cell cycle arrest, organic processes, and apoptosis in response to cellular stress. Previous ChIP and expression studies of individual genes suggest that p53 sites in HERV LTRs may be part of the p53 transcription program and directly regulate p53 target genes in humans. Therefore, the direct interaction between p53 and the LTRs of HML-2 may actively participate in gene regulations, thereby contributing to the physiological and pathological functions of LTRs of HERVs.
Genome-wide DNA hypomethylation might lead to HERV expression in cancers. The HERV-K_22q11.23 provirus is strongly expressed in prostate cancer and a spliced accessory Np9 transcript in some tumors [42]. DNA methylation in the LTR is decreased. HERV-K17 expression is significantly diminished in prostate cancer, independently from LTR methylation. The expression of these HERVs is observed in androgen-responsive prostate cancer cell lines. Their LTRs sequences contain steroid hormone-responsive elements, which bind the androgen receptor and demonstrate androgen responsiveness. The HERV-K activities in prostate cancer is specific, because LINE-1 hypomethylation does not lead to generalized overexpression, suggesting significant roles in prostate carcinogenesis. HERV-E 4.1 env RNA, HERV-H pol RNA, and HERV-W gag RNA are also detected in prostate cancer epithelial cell lines [43].
Plasma from prostate cancer patients exhibits the humoral response against epitopes of HERV-K and HERV-H envelop proteins but not against HERV-W [44]. HERV-H is also implicated in prostate cancer. HERV-K gag protein is frequently detected in prostate tissues, and the expression is regulated by methylation of the promoter region and androgen stimulation [45]. Antibodies reacting with HERV-K gag protein are more frequently detected in sera from prostate cancer patients than in healthy individuals. The presence of serum antibodies is correlated with a poor prognosis of prostate cancer. HERV-K gag RNA and protein expression are increased in malignant regions of the prostate in men with prostate cancer protein [43].
The locus-specific HERV expression is reported in different types of cancers [46]. There are active HERV elements and differentially expressed HERVs in prostate, breast, and colon cancer. Differentially expressed host genes in these cancers include genes involved in demethylation and antiviral response pathways. This result indicates the pathogenic mechanisms of HERVs in cancer development. A subset of differentially expressed HERVs are also identified as differentially expressed genes in those 3 different cancers. One hundred fifty-five HERVs are differentially expressed in all 3 cancer types, and patterns in HERV expression are specifically identified. These can be used to reveal potential biomarkers and therapeutic targets in these cancers.
Retrovirus human immunodeficiency virus (HIV-1) is detected within testicular germ cells (TGCs) in the testis from an infected human [47]. HIV-1 enters TGCs exposed to infected lymphocytes in the testis from an HIV-positive man, and then is integrated to express the early protein. The study shows that TGCs can support the entry and integration of retrovirus, which represents a way to incorporate into human germ line, as happened for HERVs.
Urothelial Carcinoma
HERV-H LTR-associating protein 2 (HHLA2) is highly expressed in multiple solid malignant tumors, including bladder urothelial carcinoma (BUC) [48]. The HHLA2 expression level is significantly associated with tumor size, stage, grade, and lymph node metastasis, and is an independent prognostic factor of tumor metastasis. High HHLA2 expression is significantly correlated with a poor prognosis of BUC. Therefore, HHLA2 is likely to independently predict unfavorable prognosis and serve as a potential diagnostic marker for BUC. Comparison of HERV transcription profiles in bladder cancers reveals at least 6 ubiquitously active HERV subgroups: E4-1, HERV-R, ERV9, HERV-K-T47D, NMWV3, HERV-KC4 [49]. Although the transcription pattern is primarily similar in between human urothelial carcinoma, nonmalignant urothelial tissue, 4 tumorderived cell lines, and a nonmalignant urothelial cell line, activities of HERV-E4-1, HERV-K (HML-6), and HERV-T (S71- TK1) tend to be lower in carcinoma. Six ERV-E4-1 loci are differentially regulated in urothelial carcinoma cells compared to normal tissue. HERV-Ec1 and HERV-Ec6 are 2 full-length proviruses in introns of PLA2G4A and RNGTT in antisense orientation, respectively. PLA2G4A encodes a phospholipase A2 and RNGTT, a bifunctional mRNA-capping enzyme exhibiting both RNA guanylyltransferase and RNA 5’-phosphatase. PLA- 2G4A is dysregulated in various tumors, and RNGTT is associated with cecum cancers. Significantly, the transcription of PLA2G4A and HERV-Ec1 is antagonistically regulated each other in human urothelial cells; thus HERV-Ec1 may contribute to fine-tuning of PLA2 expression and urothelial tumorigenesis.
It is reported that smoking may induce HERV transcription in the human urothelium [50]. HERV-E4-1, HERV-T S71-TK1, and HERV-K HML-6 transcripts tend to increase in human epithelium from smokers and human dermal fibroblasts treated with smoker’s urine [51].
Clear Cell Renal Cell Carcinoma
Computational workflow identifies more than 3,000 transcriptionally active HERVs within The Cancer Genome Atlas pancancer RNA-Seq database and indicates that HERV expression is significantly associated with clinical prognosis in several tumors, including clear cell renal cell carcinoma (ccRCC) [52]. HERV signatures associated with RIG-I-like signaling and retroviral antigen activation of adaptive immunity can predict ccRCC patient survival independent of clinical stages and molecular subtypes [51]. Potential ccRCC-specific HERV epitopes identified in ccRCC ribosome profiling dataset bind HLA in vitro, and MHC tetramer-positive T cells to the predicted epitopes are detected. In addition, those identified HERV sequences are highly expressed in ccRCC tumors that are responsive to inhibition of programmed death receptor 1. HERV expression could be a biomarker for patient prognosis and response to immunotherapy [53]. HERVs may activate the IFN signaling pathway by a viral mimicry process to enhance antitumor immune responses.
HERVs IN KIDNEY
Eyes absent homolog 1 (EYA1) is a TF that plays a role in the development of the eye, ear, branchial arches, and kidney [54]. The human EYA1 gene is located in 8q13.3 region of chromosome 8, and mutation in EYA1 is the most common cause of branchio-oto-renal and branchio-otic syndromes, characterized by preauricular pits, hearing loss, and branchial and kidney defects. Significant chromosomal aberrations of 8q13, including complex rearrangements, occur in about 20% of affected individuals, but there are also rare cases of microdeletions involving the EYA1 gene. The submicroscopic deletion can be mediated by 2 HERVs [55]. The DNA sequence analysis shows that the distal and proximal deletion breakpoints perfectly map within the 107 bp homologous sequences in the LTRs of HERV1 elements on both sides, suggesting nonallelic intrachromosomal homologous recombination.
Pigs are utilized as potential donors for xenotransplantation [56,57]. It is shown that porcine ERV (PERV) can infect both human and other primate cells in vitro, presenting an ERV transmission in xenotransplantation [58,59]. The PERV originated from porcine donor cells is detected in rhesus monkey (Macaca mulatta) bladders in xenotransplantation of the kidney and ureter [60]. Porcine cell-microchimerism was responsible for the detected PERV DNA and RNA, but the PERV genomic sequence was not integrated into the recipient chromosome.
REPRODUCTION
Germ Cells and Early Embryos
HERV-K (HML-2) was expanded over the last 5-20 million years ago [32]. HERV-K-related sequences are also present in the Old World monkeys. Still, the most recent Hominidae-specific HERV-K elements contain the LTR5Hs regulatory sequences, which may drive viral protein expression in naive human embryonic stem cells (hESCs) [61,62]. The provirus expression may stimulate a viral restriction factor IFITM-1, promoting immunological protection against reinfection of HERV-K-like retrovirus in early human embryos before implantation. Including LTR5H elements, other HERV elements such as SINEVNTR- Alu (SVA) and HERV-H-associated LTR7 are marked by H3K27ac in human pluripotent cells [63]. Also, some KLF family TFs crucial for pluripotency bind and activate TE Embedded eNhancers (TEENhancers) present in LTR5Hs and SVA elements to facilitate human embryonic genome activation. Therefore, the epigenetic-sensitivity and active gene-regulating property of the evolutionarily young Hominidae-specific TE elements in HERVs have functioned in pluripotency and early human development.
HERVs contribute to the proper formation of human primordial germ cells (hPGCs), which founders germline cells that give rise to oocytes or sperm in mature humans [64]. Some specific transcriptional networks are activated during the hPGC specification, which happens at day 11–12 postfertilization right before gastrulation. For example, various TFs such as NANOG, PRDM1, TFAP2C, and PRDM14 play critical roles in hPGC specification and maintenance [65-71]. Although these networks are largely conserved in mouse primordial germ cell (PGC) development, there is indeed some human-specific feature [62]. For example, a naive enhancer at the POU5F1 (OCT4) locus in hPGCs is utilized, which is not conserved in mice.
However, mouse model studies still provide tremendous insights to help understand the cellular and molecular mechanisms of ERV functions. DNA methylation protects male germ cells from transposon activity. The DNA methyltransferase DNMT3C protects male germ cells from transposon activity [72]. m6 A RNA methylation restricts ERVs by targeting 5´-untranslated region of ERV-derived elements in mice [73]. The complex of methyltransferase-like METTL3 catalyzes methylation of ERV mRNAs–METTL144 proteins, and depletion of METTL3–METTL14, along with their accessory subunits WTAP and ZC3H13, leads to increased mRNA abundance of intracisternal A-particles (IAPs) and related ERV-K elements specifically. IAP mRNA and protein abundance are dynamically and inversely correlated with m6 A catalysis. m6 A methylation reduces the half-life of IAP mRNA, which occurs by the recruitment of the YTHDF family of m6 A reader proteins. RNA methylation protects cellular integrity by clearing reactive ERV-derived RNA species, which can trigger inflammatory responses, as seen in human diseases [74].
HERVs are dynamically expressed during early embryogenesis and differentiation [75]. Although several HERV subfamilies are expressed in a tight spatiotemporal mode in early human embryos, HERV expression is largely repressed in early human embryos, and its derepression is associated with lethality at various embryonic stages [30,76]. It has been reported that ERV derepression triggered upon acute perturbation of the repression machinery generates ERV RNAs at hundreds or maybe even thousands of genomic loci, disrupting phase-separated transcriptional condensates enriched in RNA polymerase II in the nucleus [12].
Piwi-interacting RNAs (piRNAs) are predominantly expressed in germ cells and function in gametogenesis in various species [77-84]. Disruption of any piwi paralogs in Drosophila or zebrafish results in infertility in both males and females [85- 89], but interestingly in mice, only in males [90-96]. Piwi-deficient female mice are fertile, and mouse oocytes express species- specific expression of endo-siRNAs, but relatively low piRNAs [97-99]. Mice genome also lacks a piwi paralog piwil3, which is common in most mammals, including humans [77,79]. However, the functional piRNA pathway is essential in male and female fertility in golden hamsters having all 4 piwi genes [79,100]. Golden hamsters with impaired piRNA pathways exhibit defective embryogenesis, reproductive development, or spermatogenesis, and produce infertile oocytes [100- 102]. Interestingly, the study using Piwil1-deficient golden hamsters generated by CRISPR-Cas9 shows that PIWIL1-piRNAs function in silencing ERVs in the oocytes and subsequent impairment of maternal mRNA degradation and zygotic genome activation in the early embryos [100,103]. PIWIL1 is required to generate ERV-related PIWI3 piRNAs, because the ERV-related PIWIL3 piRNAs are nearly absent in Piwil1-deficient oocytes and maternal Piwil1-deficient embryos.
Ten-eleven translocation (Tet) methylcytosine dioxygenases participate in DNA demethylation, playing a crucial role in stem cell pluripotency and differentiation by DNA methylation patterning [104-106]. Tet1-mediated 5hmC signals are critical in the DNA demethylation process during PGC development and meiosis [107-109]. Tet1 deficiency causes insufficient demethylation, leading to the failure of meiotic gene activation that results in meiotic defects such as impaired homologous pairing and recombination. Therefore, Tet1-deficient mice show a reduced number of oocytes and reduced follicle reserve, which are significant symptoms of premature ovarian failure (POF) syndrome [110]. Tet2 deficiency accelerates reproductive aging by delaying meiotic progression and reducing oocyte quality in adult female mice but does not reduce the number of oocytes [111]. The single cell transcriptome data of oocytes suggest that Tet1 deficiency alter ERVs, elevate organelle fission, upregulate signaling pathways for Alzheimer diseases, and downregulate X-chromosome-linked genes such as Fmr1, whereas Tet2 deficiency reduce DNA repair, meiosis progression, the spindle assembly checkpoint, and actin cytoskeleton dynamics, and alters chromosome alignment and segregation. Therefore, Tet1 enzyme plays a role in maintaining oocyte quality, oocyte number, and follicle reserve, while Tet2 enzyme in oocyte development and quality assurance.
Male Reproduction
Sperm
Mammalian spermatozoa can bind and internalize exogenous DNAs into their nucleus [112,113]. Integration of a small portion of the internalized DNA happens at specific sites in the sperm genome, probably highly enriched with undermethylated sequences, many of which correspond to retrotransposons where endogenous retrotransposition events occur [114,115]. Mouse spermatozoa are active in transcription, splicing, and reverse transcription [116]. Exogenous retroelements, including HERV-K10, can be incorporated into the mouse sperm genome and transferred into the mouse oocyte, causing genetic alterations in embryos [117]. Finally, HERV-K10, as well as other retroelements such as LINE-1 and SVA, is expressed in ejaculated human spermatozoa, which are capable of integrating cloned active retrotransposons into their genome and mediating active transcription. Endogenous retrotransposition is the event that occurs in the very early development of human embryos and is normally controlled by pi-RNAs and methylation [118,119]. Impaired control of the human retroelements may negatively affect cellular proliferation, genome stability and methylation, and preimplantation embryo development [120]. Therefore, extracellularly introduced retroelements can give rise to effective retrotransposition events, which cause somatic cell phenotype changes and interfering with differentiation and propagation.
Rats homozygous at the hypodactyly (hd) locus (hd/hd) show impaired limb development and spermatogenesis [121]. Sperm from the mutant rats exhibit tail fragility and decapitated heads, which are symptoms of human teratoasthernospermia. An ERV-K8e family member is inserted into intron 10 of the centrobin (Cntrob) gene in the hd locus, disrupting the normal splicing of Cntrob transcripts [122]. Knockdown of centrobin inhibits centriole duplication, resulting in microtubule assembly defects and abnormal nuclear morphology. Centrobin interacts with keratin 5-containing intermediate filaments at the marginal ring of the spermatid acroplaxome. In the hd mutant spermatids, the acroplasome marginal ring is abnormal, and the centrosome is separated from its normal attachment site of the nucleus. These phenotypes correlate with a disruption of the head-tail coupling apparatus, which can lead to spermatid decapitation and tail fragility. Centrobin is also known as Lip8 (LYST [lysosomal trafficking regulator]-interacting protein 8) and Nip2 (Nek2-interacting protein 2) [123,124].
Y chromosome
Azoospermia factor region (AZF) in the long arm of the human Y chromosome contains many functional genes and transcription units essential for spermatogenesis [125]. AZFa, AZFb, and AZFc are the 3 key subregions, and there may be a fourth subregion, AZFd. Microdeletion in the AZF regions is one of the major genetic factors in male infertility. Many of the microdeletion events result from nonreciprocal intrachromosomal recombination between homologous sequences, which are located in repeated regions. For example, AZFa deletions can happen through nonreciprocal homologous recombination between 2 HERV sequences [126,127].
HERVs are also associated with testis-specific transcripts linked to the Ys (TTYs) in the AZFb region [128,129]. Computational screening reveals that TTY9, 10, and 13 are regulated by HERVs in the AZFb region. Homologous recombination between LTRs of the TTY13-associated HERV-K14C can make a microdeletion in TTY13 gene, and these deletions are more frequently observed in males with azoospermia or oligozoospermia than in fertile ones. Therefore, HERV-driven genomic change in the AZFb region may contribute to some types of human idiopathic male infertility. The finding also supports that TTY transcripts play a critical role in spermatogenesis and male fertility.
HERVs are accumulated with density in the male-specific region of the Y chromosome (MSY), and HERV-K14C seems to preferentially into the human Y chromosome [130]. The human genome contains 146 copies of HERV-K14C, and 29 of them are located in the Y chromosome, mostly dispersed in the palindromic region. Three distinct HERV-K14C-related transcripts are exclusively expressed in human testis tissue. Phylogenetic analysis of the LTRs indicates that all HERV-K14C sequences in the Y chromosome are generated as pairs of identical sequences. The Y chromosome amplicons existed in our common ancestors, and the duplicated pairs arose after the divergence of great apes approximately 8–10 million years ago. Therefore, HERV-K14C sequences have contributed to genomic diversification of the Y chromosome during the speciation of the great ape lineage.
Female Reproduction
Placenta
HERVs play a critical role in mammalian placental and embryonic tissue development, modulating endocrine activities and cell fusion events in the trophoblast cells. Pleiotrophin, a human growth factor expressed in trophoblast, is controlled by the HERV LTR promoter [131]. Syncytin-1 (SYN-1) and syncytin-2 (SYN-2) are products of HERV env genes, HERV-W1 and HERV-FRD, respectively [2,3,11]. Syn-1 and Syn-2 bind their respective receptors, ASCT-2 (sodium-dependent neutral amino acid transporter type 2, aka solute-linked carrier family A member 5 [SLC1A5]) and domain-containing protein 2a (MFSD2a) in human trophoblasts, inducing fusion of individual trophoblast cells into a multinucleated layer of syncytiotrophoblasts, which form an outer layer of placental villi and in direct contact with maternal blood flow [51]. Suppressyn (SUPYN) is a truncated ENV protein encoded by the HERVH48-1 env gene from a HERV-H family virus, which interferes with cell-cell fusion [132,133]. ERVMER34-1 is another HERV-derived protein, which is also expected to negatively regulate cell fusion in cytotrophoblast [134]. SUPYN shares the common receptor SLC1A5 with SYN-1. SLC1A5 is broadly expressed in placental and decidual cells, whereas the expression of SYN-1 and SUPYN are mostly confined in trophoblast cells.
Endometria
In addition to their well-known roles in placental and embryonic tissues, it has also been questioned whether HERVs play a role in endometrial homeostasis before implantation. Several studies suggest that HERVs are involved in endometrial cancer and endometriosis [135-137]. It has been demonstrated that the endometrium isolated from some women with recurrent pregnancy loss (RPL) contains substantially more transcripts of the retroviral HERV-K GAG gene than controls who have had at least one successful pregnancy [138]. Although the study was preliminary, the correlation suggests that high HERV expression may induce endometrial inflammatory reactions, which may act concurrent with other factors that can result in endometrial dysfunction, such as viral infections, genetic variations, and environmental influences [139,140]. Notably, dysregulated expression of HERVs demonstrated in RPL is possibly associated with the RPL-linked polymorphisms of methylenetetrahydrofolate reductase, a methyl donor-producing enzyme required for the proper functional cellular methylation pathways crucial for DNA methylation. Considering the vital impact of ERVs in endometrial health and preimplantation development of embryos, ERV regulation is likely to be critical for successful embryo implantation following in-vitro fertilization [141].
CONCLUSION
Since infected human germline cells and incorporated into the human genome, the mostly inactive HERVs have actively participated in human biology. Fine-tuning HERV repression is essential in hPGCs and hESCs. Altered HERV expressions are broadly observed in various urological cancers, including prostate and urothelial cancers. While specific envelop-gene-derived proteins, such as SYNs and SUPYN, function in the female reproductive procedure of placental development, HERVs also significantly affect male reproduction quality by controlling male-specific gene expressions. HERV transcript profiles are potential biomarkers for prostate cancer diagnosis and possibly for other disorders exhibiting altered HERV expressions. Understanding HERV features and functions in urological subjects will help develop practical applications to be utilized for diagnosis and prognosis evaluations in urology clinics.
Notes
Funding/Grant Support
This research was supported by the Basic Science Research Program through the National Research Foundation of Korea (NRF) funded by the Ministry of Education, Science and Technology (2021R1F1A1049211).
Conflict of Interest
No potential conflict of interest relevant to this article was reported.
AUTHOR CONTRIBUTION STATEMENT
·Conceptualization: SL, JA
·Funding acquisition: JA
·Methodology: SHK
·Project administration: SL, JA
·Visualization: SHK
·Writing - original draft: SL
·Writing - review & editing: SL, SHK, JA
