Extracellular Vesicles as a Source of Urological Biomarkers: Lessons Learned From Advances and Challenges in Clinical Applications to Major Diseases
Article information
Abstract
Extracellular vesicles (EVs) not only eliminate unwanted molecular components, but also carry molecular cargo essential for specific intercellular communication mechanisms. As the molecular characteristics and biogenetical mechanisms of heterogeneous EVs are different, many studies have attempted to purify and characterize EVs. In particular, exosomal molecules, including proteins, lipids, and nucleic acids, have been suggested as disease biomarkers or therapeutic targets in various diseases. However, several unresolved issues and challenges remain despite these promising results, including source variability before the isolation of exosomes from body fluids, the contamination of proteins during isolation, and methodological issues related to the purification of exosomes. This paper reviews the general characteristics of EVs, particularly microvesicles and exosomes, along with their physiological roles and contribution to the pathogenesis of major diseases, several widely used methods to isolate exosomes, and challenges in the development of disease biomarkers using the molecular contents of EVs isolated from body fluids.
INTRODUCTION
In multicellular organisms, cells communicate through a myriad of tightly regulated molecules, and intercellular communication systems have evolved with high efficiency for adaptation and biodiversity. Intercellular communication occurs at numerous levels between cells, among cells and tissues, and across organisms. Among these, the secretion and uptake of molecular contents, including proteins, lipids, metabolites, and nucleic acids, from adjacent or remote cells is an efficient system that modulates cellular homeostasis and adaptation. The amounts and types of secreted molecules are altered during development and aging, and are determined by specific stimuli (e.g., drugs) or pathological conditions. In several major diseases, the secretion of extracellular molecules may contribute to disease development or progression in a disease-specific manner. Therefore, secreted molecules are considered as both candidate diagnostic biomarkers and therapeutic targets. To avoid degradation by extracellular enzymes and ensure circulation throughout the body, the compartmental segregation of secreted molecules (e.g., by membranous vesicles) is required. Exosomes and microvesicles (also known as ectosomes or shedding vesicles) are extracellular membranous compartments containing a variety of cargo molecules and are generated via tightly regulated exocytosis and membranous budding or shedding, respectively. Such extracellular vesicles (EVs) can be readily found in most body fluids, such as serum, plasma, urine, and cerebrospinal fluid (CSF), which enables the molecular contents of EVs to be analyzed as disease biomarkers. Notably, recent studies have provided evidence that exosomes are the major EVs involved in intercellular communication via the autocrine, paracrine, and circulatory systems. This review summarizes the characteristics of EVs, their fates from biogenesis to secretion, and their physiological and pathological roles. Regarding the methodological aspects of exosome purification from body fluids for biomarker development, we discuss commonly used methods for exosome isolation, their advantages and disadvantages, and current challenges in exosomal biomarker development.
EXOSOMES AND MICROVESICLES: MOBILE INTERCELLULAR COMMUNICATORS
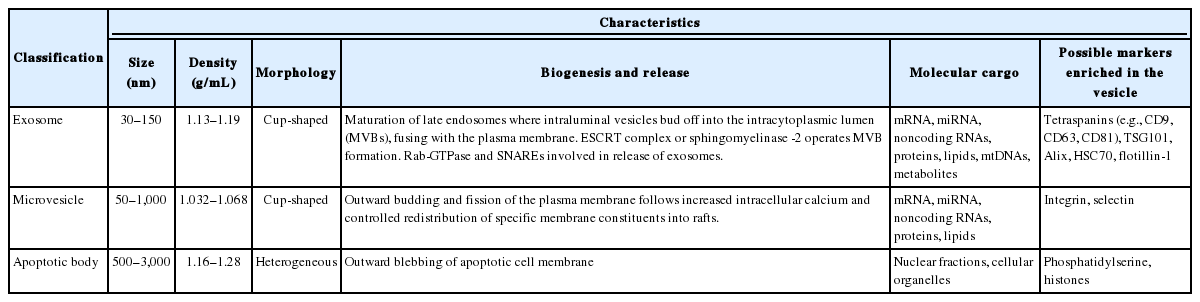
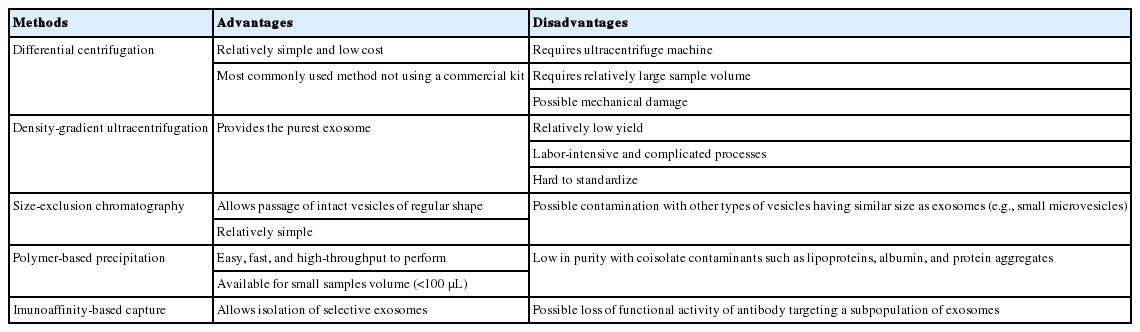
EVs are released from most types of cells, including hematopoietic cells, astrocytes, neurons, microglia, epithelial cells, endothelial cells, myocytes, and stem cells [1,2]. EVs comprise microvesicles, exosomes, and apoptotic bodies. Microvesicles and exosomes originate from healthy or stimulated cells, and are smaller in size than the apoptotic bodies released from dying cells (0.5–3 μm). More importantly, the molecular cargo and biogenic origins of EVs are quite variable. Table 1 summarizes the characteristics of EVs. The molecular characteristics of secreted EVs vary according to the type of parent cell and can determine the functions of EVs in mobile intercellular communication. Although the specificity of EVs is presumably determined by protein-protein interactions between membranous proteins and certain target cell types, the mechanisms regulating the mobility of EVs and the assembly of luminal constituents remain largely unknown. It has been widely accepted that cells subjected to stimuli such as inflammation, hypoxia, and metabolic deregulation (e.g., insulin resistance) exhibit increased release of EVs, particularly exosomes, compared to unstimulated cells. Therefore, under certain pathogenic conditions, the molecular contents and secretion amounts of EVs may determine whether EVs play roles in compensatory homeostasis against pathogenic mechanisms or in the propagation of pathologies. For example, pancreatic cells subjected to apoptotic stimuli release EVs that stimulate cell proliferation to maintain homeostasis. In contrast, neuronal cells stimulated by proteinopathic stimuli (e.g., α-synuclein [α-syn] in Parkinson disease [PD]) release exosomes that transfer pathogenic proteins (synucleinopathy) through neuron-to-neuron transmission [3-5]. Therefore, the mechanisms of biogenesis, release, and uptake of EVs under certain pathologic conditions must be elucidated in order to understand disease progression and to identify disease biomarkers and therapeutic targets.
Biogenesis and Secretion of EVs
Although the size and density ranges of exosomes and microvesicles may overlap, their biogenic processes are distinct (Fig. 1). During stress or maintenance of homeostasis, microvesicles are formed and shed at the plasma membrane, which is enriched in phosphatidylserine (PS) in the outer membrane leaflet, with a lipid asymmetry caused by PS externalization. At first glance, the release of microvesicles seems to be a simple process of outward budding of the plasma membrane followed by a fission event. However, energy-dependent mechanisms such as membrane curvature [6], asymmetric PS movement by aminophospholipid translocase [7,8], cytoskeleton involvement [9], and redistribution of molecular cargos [10] are involved in microvesicle biogenesis. Since not all proteins are incorporated into microvesicles, a variety of proteins without signal peptides are selectively transported via the ARF-6-mediated endosomal recycling pathway, which is an unconventional protein secretory pathway that discriminates between secreted proteins with or without signal peptides. In addition, nucleic acids and lipids are incorporated into microvesicles and transferred to recipient cells.
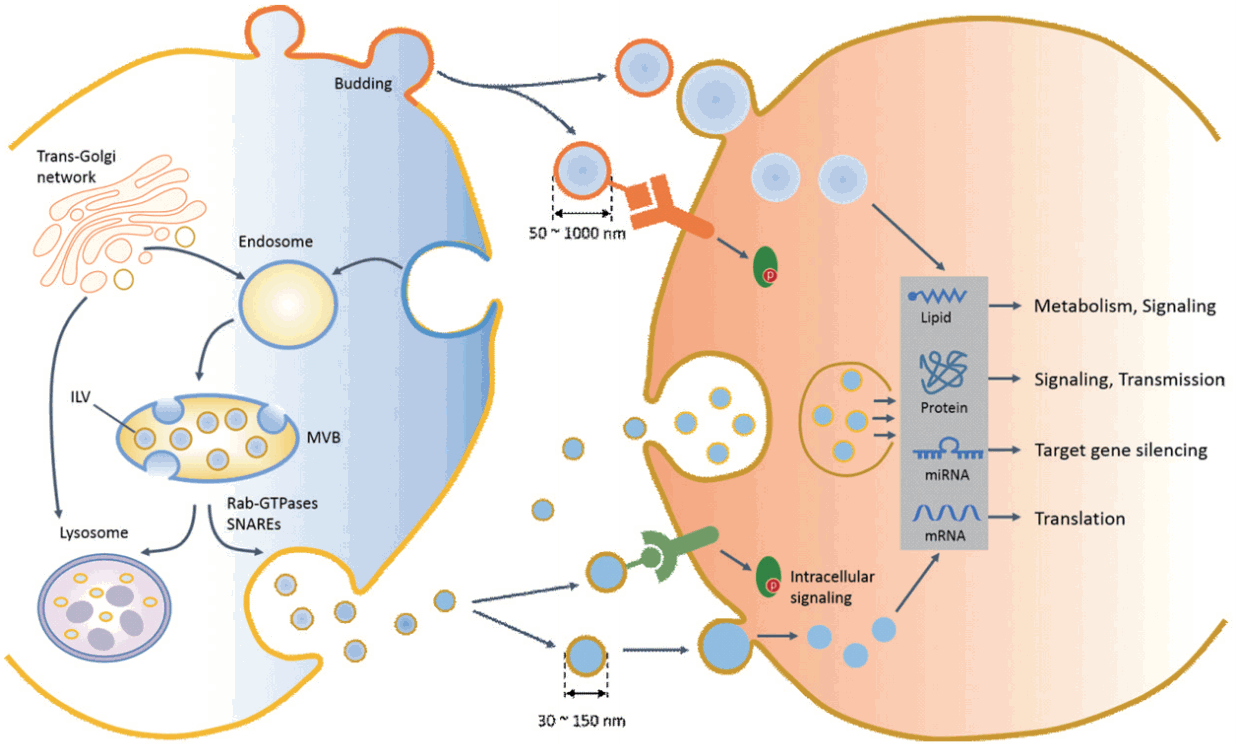
Biogenesis, release, uptake, and cellular functions of exosome and microvesicles in recipient cells. An early endosome matures and forms a multivesicular body (MVB) containing an intraluminal vesicle (ILV), which buds off into the lumen of the late endosome. The MVB then directly fuses with the plasma membrane (exosome release) or with the lysosome (degradation of intraluminal content). The endosomal sorting complex required for transport (endosomal sorting complex required for transport) or ceramide contributes to processes of MVB formation, protein cargo sorting, and lysosomal degradation or release of the exosome. Multiple mechanisms and factors, including intracellular calcium changes, intracellular lipids, the small Rab family GTPases, and the soluble N-ethylmaleimide-sensitive factor attachment protein receptor (SNARE) system, are involved in the transport and fusion of MVBs with the plasma membrane (exosome release). Microvesicles are produced by the direct outward budding of plasma membrane. After release, extracellular vesicles (EVs) interact with target cells via several ways, including interaction of an EV ligand with a protein (receptor) on the target cell membrane, endocytosis, phagocytosis, or direct delivery of EV cargo into the cytosol by membrane fusion. The molecular cargos of EVs regulate diverse cellular functions of target cells through intracellular signaling, gene regulation, and metabolism. miRNA, microRNA.
Although the mechanisms for exosome and microvesicle biogenesis may share mechanistic elements, exosomes have an endocytic origin and are released via the fusion of multivesicular bodies (MVBs) with the plasma membrane (exocytosis). In late endosomes, MVB formation is tightly regulated by a piece of sequential molecular machinery known as the endosomal sorting complex required for transport (ESCRT) complex on the endosomal membrane [11]. Alternatively, an ESCRT-independent pathway involving ceramide generating sphingomyelinase was recently shown to be involved in intraluminal vesicle (ILV) formation and the release of exosomes in tumor, glia, and antigen-presenting cells [12,13]. Exosomes are enriched with lipid raft-associated molecules such as cholesterol, sphingolipids, glycerophospholipids, and prostaglandins. In addition to protein and lipid incorporation, exosomes contain genomic materials such as messenger RNA (mRNA), microRNA (miRNA), and mitochondrial DNA. Although Argonaute 2 and the miRNA-induced silencing complex capture miRNAs during early endosome formation, more than 90% of Ago protein-bound miRNAs are not found in EVs. Therefore, it remains to be elucidated how miRNAs are selectively incorporated into ILVs before exosome release. It has been widely accepted that ESCRT and sphingomyelinase are critical for the generation of MVBs, which subsequently fuse either with lysosomes for protein degradation or with the plasma membrane for exosome release.
The distinct fates of MVBs (i.e., lysosomal degradation versus exosome release) may depend on the composition of MVBs, which is tightly regulated by molecular machinery [14]. Exosome release processes require multiple mechanisms, involving whether MVBs fuse with lysosomes or the plasma membrane, the mobilization of MVBs, and docking and fusion with the plasma membrane. Although these mechanisms are not fully understood, the fate of MVBs is dependent on the degree of cholesterol or lysobisphosphatidic acid enrichment [15] in MVBs. Several molecular switches of the Rab GTPase family were the first mechanisms identified in exosome secretion. Recent studies have provided evidence that the Rab11, Rab27, and Rab35 pathways are clearly involved in exosome release [16-19]. For example, Rab27a and Rab27b play key roles in exosome secretion by contributing to MVB trafficking and docking with the plasma membrane [18]. Rab11 is also involved in the fusion of MVBs with the plasma membrane in a calcium-dependent manner [19]. Membrane fusion of MVBs to the plasma membrane is regulated by the soluble N-ethylmaleimide-sensitive factor attachment protein receptor (SNARE) complex, which is involved in the calcium-regulated exocytosis of conventional lysosomes. The v-SNAREs (also referred to as R-SNAREs) anchored into the MVB membrane combine with t-SNAREs (also referred to as Q-SNAREs) located in the plasma membrane to form a membrane-bridging SNARE complex, which results in the exocytosis of vesicles in MVBs (exosomes).
Physiological Roles of EVs
Secreted EVs selectively bind to recipient cells in a target cell-specific and condition-dependent manner. Adhesion molecules, such as integrins expressed in the exosomal membrane, may determine target cell specificity. Large-scale protein analysis using proteomic techniques has revealed that exosomes do not contain a random set of intracellular proteins, but rather a specific array of very limited proteins from intracellular compartments. Exosomal proteins have been compiled in the Exo-Carta compendium [20]. In addition, the presence of genetic information, including mRNAs and miRNAs, in EVs strongly suggests that EVs transfer genetic messages regulating gene expression in a target-specific manner [21]. EVs deliver diverse molecules to recipient cells and therefore are multifunctional molecular complexes that control the fundamental and homeostatic functions of cells (Fig. 2). For example, EVs regulate immune responses by triggering adaptive immunity and suppress inflammation by regulating immune cell activity, differentiation, and proliferation [22-24]. In the brain, molecules associated with neuronal function and neurotransmission in EVs contribute to intercellular communication between neural cells (e.g., neuron-glia interaction), synaptic plasticity, and neuronal activity [25]. In particular, reciprocal communication between neurons and neighboring cells, including astrocytes, microglia, oligodendrocytes, and cells, in the blood-brain barrier via EVs is a fundamental system for maintaining not only neuronal activity, but also the microenvironment of the neural system. However, certain pathogenic molecules in EVs can play roles in the progression of neurodegenerative diseases. EVs from the central nervous system can be found in the plasma as well as CSF, which allows the molecular cargo of EVs to be used as biomarkers of neurologic diseases. EVs are also potentially powerful shuttles for the delivery of therapeutic agents, and EVs from adult mesenchymal stem cells (MSCs) can be used in regenerative medicine. The regenerative potential of the MSC-derived secretome has been recently identified and extensively reviewed [26,27]. EVs were isolated from MSCs for the first time and characterized in 2010 [28]. Although it remains unknown whether EVs released from MSCs are responsible for the pluripotent features of MSCs, the microenvironment of MSCs may be controlled by the molecular contents of EVs, such as exosomal miRNAs.
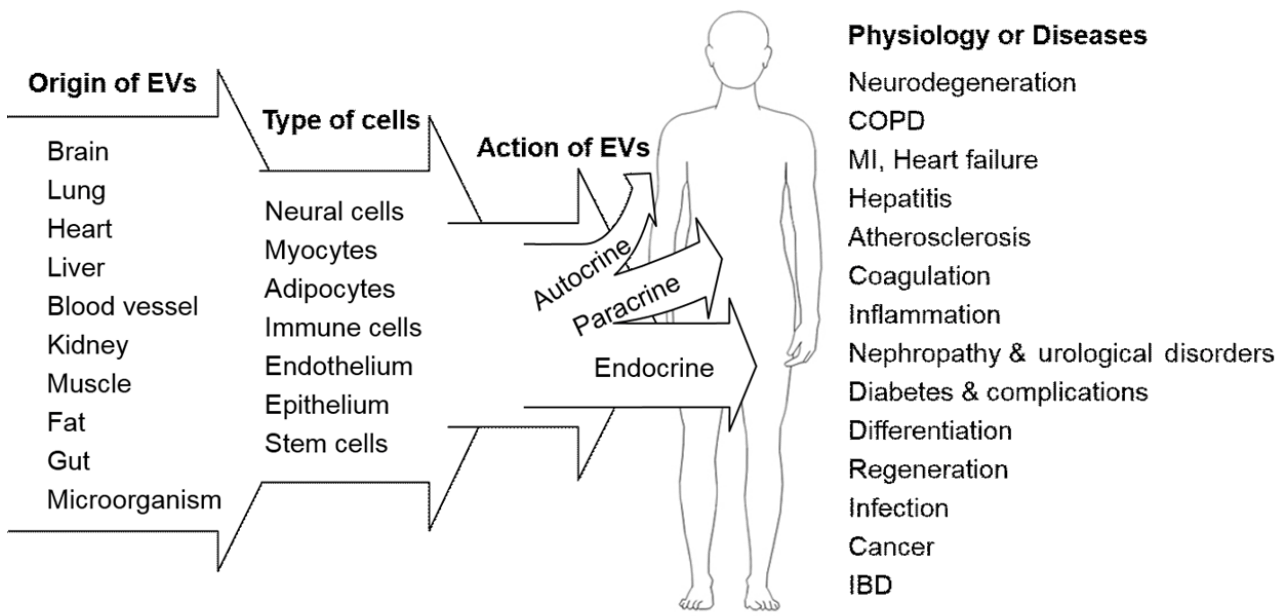
The extracellular vesicle (EV) as a multifunctional effector regulating normal cellular function or disease pathogenesis. EVs originate from most tissues and are comprised of various types of cells, which communicate with adjacent (autocrine and paracrine effects) or remote (circulating factor) cells. In addition to regulation of the physiological function of adjacent or remote cells, intercellular communication via circulating EVs presumably contributes to the development and progression of disease, remission of disease, or the protection of tissue against injury. Therefore, EVs and their molecular contents in body fluids may be candidate disease biomarkers or therapeutic targets. COPD, chronic obstructive pulmonary disease; MI, myocardial infarction; IBD, inflammatory bowel disease.
EVs in Diseases
Cancer and the prostasome
Tumor-secreted EVs transfer oncogenic molecules to neighboring cells and their local microenvironment, suggesting that EVs act as critical messengers in tumor initiation, progression, and metastasis. EVs reflect diverse changes in cell and tissue microenvironments during cancer development. Regarding tumor initiation, accumulating evidence from in vitro studies has suggested that EVs from malignant cells are involved in the malignant transformation of adjacent normal cells. For example, the uptake of epidermal growth factor receptor in tumor cell-derived EVs by endothelial cells increases the expression of vascular endothelial growth factor signaling pathways associated with tumor angiogenesis [29]. In addition, EVs from various tumor cells transfer intravesicular or membranous proteins to stimulate malignant transformation [30,31], cell proliferation signaling pathways [32], or oncogene amplification [33]. Based on in vitro studies, the role of oncogenic contents derived from exosomes in cancer has been established [34]. The association of EVs with cancer development and progression has also been tested in animal models and humans. In fact, tumor-bearing animals and cancer patients have higher levels of EVs in body fluids such as blood, urine, and saliva [35-37]. In particular, increasing evidence shows that exosomes lead to tumor progression and metastasis by increasing tumor cell migration/invasion, contributing to the formation of a metastatic niche, and remodeling of the extracellular matrix through exosomal molecules such as miRNAs or proteins [38]. Exosomal miRNAs in urologic cancer cells have been recently investigated. For example, miR-21-5p in urinary EVs was suggested as a novel biomarker of urothelial carcinoma [39]. Another study by Royo et al. revealed that cadherin 3, type 1 in urinary EVs was negatively regulated in prostate cancer, and that it may serve as a noninvasive tool to obtain information about the molecular alterations that take place in prostate cancer [40]. The upregulation of exosomal miRNAs in cancer stem cells is also associated with cancer cell proliferation and migration [41]. Recently, attempts have been made to substitute biomarkers less specific to certain types of cancer (e.g., prostate cancer antigen for prostate cancer) with exosomal miRNA profiles. Mitchell et al. [42] reported that miRNA-141 levels were increased in the serum of metastatic prostate cancer patients, which has been reproduced in other studies [43,44]. In addition, deregulated miRNAs are associated with the progression of disease and lymph node metastasis [45]. In prostate cancer, the quantification of circulating prostate EVs using nanoscale flow cytometry technology and liquid biopsy may have significant prognostic potential and, therefore, clinical utility [46]. Collectively, exosomal or EV contents (miRNAs and proteins) have been highlighted as new diagnostic and prognostic markers of cancer, although more convincing evidence in large clinical cohorts is still required. In particular, since urinary EVs might be good candidates for noninvasive sources of biomarkers of genitourinary diseases, EV-related research can be considered to be a rapidly emerging field. In addition, several obstacles to validating exosomal contents as therapeutic targets and in the isolation of exosomes from body fluids obtained by liquid biopsy should be addressed.
Metabolic disease
Metabolic homeostasis is maintained by crosstalk through hormones and metabolites among metabolic tissues, including adipose tissue, the liver, and skeletal muscle, and imbalances in these processes lead to metabolic disorders. Recently, accumulating evidence has suggested that EVs are implicated in metabolic disturbances. White visceral adipose tissue (WAT) is an active endocrine organ that regulates whole-body metabolism and metabolic signaling, as well as modulating the function of remote tissues through the secretion of adipokines (e.g., adiponectin, leptin, and resistin) [47]. In addition, WAT from obese humans releases adipokines implicated in inflammation, such as interleukin-6 and tumor necrosis factor-α, which act as key drivers of obesity-associated metabolic disease [48]. Adipokines such as adiponectin, interleukin-6, monocyte chemoattractant protein-1, macrophage migration inhibitory factor, retinolbinding protein 4, and resistin have been identified in human WAT-derived exosomes [49]. Further, exosomes derived from the WAT of obese patients were shown to inhibit insulin signaling in liver and muscle cells. In another study providing evidence for the role of WAT exosomes in metabolic diseases, an increased level of adipocyte protein 2 in blood exosomes was shown to be associated with the development of type 2 diabetes and metabolic syndrome [50]. In contrast, adiponectin in circulating EVs was found to be decreased in mice fed high-fat diets compared with lean mice [51]. Taken together, these studies suggest that adipokines in exosomes derived from WAT may play a crucial role in the development and/or progression of metabolic disorders.
Considering transcellular communication, the molecular contents of secreted WAT exosomes are transferred to immune cells, contributing to systemic inflammation and insulin resistance. In this regard, Deng et al. [52] reported that exosomes isolated from adipose tissue of obese ob/ob mice were taken up by peripheral blood monocytes of normal mice. These exosomes activated monocytes derived from bone marrow into macrophages and consequently led to the induction of insulin resistance. They also found that the exosomal molecule that induced insulin resistance was retinol-binding protein 4. Further, recent studies have provided support for the critical roles of EVs in metabolism, reporting that EVs secreted from activated macrophages were observed to impair insulin signaling in adipocytes [53,54]. However, it remains to be determined whether these systemic effects of adipose-derived exosomes also mediate communication among obese adipocytes, immune cells, and other metabolic tissues.
Neurodegenerative disorders
Several neural cell types in the brain, including neurons, glial cells, and oligodendrocytes, release EVs into the extracellular space and body fluids. EVs released from multiple types of neural cells not only contribute to intercellular communication between neural cells (e.g., neuron-glia interaction), synaptic plasticity, and neuronal activity, but are also responsible for the pathogenesis of neurological disorders [55]. Notably, several recent studies have suggested that EVs released from neural cells carry neuropathogenic molecules such as amyloid-beta (Aβ), α-syn, tau, superoxide dismutase 1, and TAR DNA-binding protein 43, which promote proteinopathic neurodegenerative diseases, as well as neuroprotective molecules against neuronal injury [2].
Parkinson disease: A hallmark of PD is the misfolding and aggregation of α-syn caused by failure of the ubiquitin-proteasome system or the autophagy-lysosome pathway. It has been reported that lysosomal inhibition increased the secretion of α-syn through exosomes [55]. Neuronal cell-derived α-syn secreted in a calcium-dependent manner is transferred to recipient cells and causes cell death, suggesting that exosomal α-syn can amplify or propagate PD pathology [56]. Recently, Shi et al. [57] reported that exogenous α-syn injected into the brains of mice was released into the blood through exosomes. Detection of exosomal α-syn has been validated in the CSF, blood, and plasma of both PD patients and heathy controls. The amount of exosomal α-syn in the plasma was shown to be higher in PD patients than in healthy controls, whereas levels of α-syn in the CSF of PD patients were lower than in healthy controls [57,58]. Another study reported lower exosomal α-syn levels in the CSF of early-stage PD patients than in neurologically normal controls, and exosomal α-syn was shown to be delivered more efficiently to cells than free α-syn, as well as to be more toxic [5,59]. More importantly, CSF exosomes isolated from PD patients were observed to induce α-syn oligomerization in vitro [59]. In addition to exosomes isolated from PD patients, exosomes in general, regardless of the amount of α-syn they contain, are known to accelerate the aggregation of α-syn due to their lipid component [60]. These results suggest that exosomes are implicated in the transmission of α-syn and/or the acceleration of α-syn aggregation, thereby contributing to the propagation of synucleinopathies such as PD and dementia with Lewy bodies. Therefore, exosomes isolated from body fluids such as CSF and- plasma may be a source of disease biomarkers, although few studies have evaluated the clinical diagnostic performance of exosomal α-syn (i.e., the presence of a diagnostic sensitivity and specificity >85%) in a large cohort [57].
Alzheimer disease: Aβ, known as the hallmark of Alzheimer disease (AD), is a fragment of amyloid precursor protein (APP) that is proteolytically cleaved by β- and γ-secretases. It was reported that β-secretase cleaves APP in early endosomes, and that Aβ is subsequently sorted into MVBs [61]. Further, APP and β- and γ-secretases were detected in exosomes derived from neuronal cells [62,63]. In clinical studies, both APP and Aβ have been shown to circulate in the CSF and plasma of human AD patients [64-66]. In addition, exosomal Aβ was found to be colocalized with other exosome markers in amyloid plaques in the brains of AD mice and in the postmortem brains of AD patients, suggesting that exosomes may be involved in the pathogenesis of AD [62,67,68]. In Aβ plaques, the aggregation of tau leading to neurofibrillary tangles has been reported to be a pathological feature of AD. in vitro studies have shown that tau secretion can be mediated by exosomes [65,69]. The uptake of tissue homogenates containing aggregated tau protein induces misfolding of tau in transgenic mice [70]. Further, exosome-related tau has been detected in human CSF samples [66]. Recently, Asai et al. [71] elegantly demonstrated that tau pathology was propagated through exosomal tau, and that microglia mediated this process. Taken together, these studies suggest that exosome-mediated secretion of Aβ and tau may play an important role in AD development and progression. Nonetheless, some studies have reported that exosomes exerted beneficial effects against the pathology of Aβ species [72-74]. Specifically, glycosphingolipids and insulin-degrading enzyme in exosomes were reported to bind to neurotoxic Aβ and to promote Aβ degradation. In the brain, several types of neural cells release exosomes containing different molecular content profiles, and thus exosomes may play different roles in physiology and pathogenesis. In addition, exosomal content profiles can be altered by genetic and microenvironmental factors that cause pathogenesis. For examples, Cheng et al. [75] found that 16 exosomal miRNAs in blood exosomes isolated from AD patients were different from those in healthy controls, and these were correlated with apolipoprotein E genotypes. In addition, Kapogiannis et al. [76] reported that the level of dysfunctionally phosphorylated type 1 insulin receptor substrate in neural-derived blood exosomes in preclinical AD patients was higher than in healthy controls. Subsequently, to evaluate the diagnostic potential of exosomal Aβ and phosphorylated tau, they performed precipitation to isolate neural-derived blood exosomes and found that the levels of p-tau and Aβ1-42 in extracts of neural-derived exosomes can be used as prognostic indicators of AD development [66]. They also examined the molecular cargo of plasma astrocyte- and neuron-derived exosomes and found that exosomal proteins were of diagnostic or prognostic value for AD [77,78]. However, it should be noted that their use of a commercially available precipitation kit for the isolation of exosomes may not be specific enough for exosome-specific biomarkers due to issues related to purity (see below). In addition, accumulating evidence suggests that miRNAs are differentially expressed in body fluids, including CSF, blood, and urine, of AD patients [79-81]. Recently, increased miRNA-34a levels were observed in the brains of AD mice and patients [82]. Using miRNA-34a-overexpressing neuronal cells, it was confirmed that miR-34a could be transferred to neighboring neurons through exosomes. In addition, 2 exosome-enriched miRNAs, miR-9-5p and miR-598, were found in reduced quantities in the CSF of AD patients compared with heathy control subjects, whereas their expression levels were higher in the exosome-enriched CSF of AD patients [83]. Therefore, under physiological or pathological conditions, the identification of various exosome-mediated pathways can provide insights into the pathogenesis of AD, diagnostic possibilities, and therapeutic strategies.
EVALUATION OF PURIFICATION METHODS FOR EXOSOMES FROM BIOLOGICAL FLUIDS
Differential Centrifugation
Differential centrifugation is the most commonly used method for exosome isolation from cell culture media and body fluids. This method involves serial centrifugation steps performed at different forces in order to remove components such as cell debris, apoptotic bodies, and microvesicles from exosomes. Primary centrifugation of culture media or a body fluid is performed at 300×g for 10 minutes to remove larger contaminants, such as dead cells and cell debris. The supernatant is then purified from the cell debris and apoptotic bodies by secondary centrifugation at 2,000×g for 20 minutes. Tertiary centrifugation at 10,000–20,000×g for 30 minutes allows for the generation of a pellet (microvesicles) on the tube wall. In some cases, this part of the protocol is modified by supernatant filtration through a 0.22-μm filter to remove particles larger than 200 nm. Finally, an exosomal pellet is generated by ultracentrifugation at 100,000×g for 2 hours, followed by washing of the pellet in phosphate-buffered saline and another round of ultracentrifugation. Importantly, the viscosity of the samples is correlated with the purity of the isolated exosomes. Thus, samples with high viscosity may require a longer ultracentrifugation step and higher centrifugation speed. In addition, viscosity is associated with the type of centrifugation rotor (i.e., fixed angle vs. swinging type) and the k-factor (pelleting efficiency) used in exosome sedimentation (i.e., swinging-bucket versus fixed-angle rotors) [84,85]. In our experience, approximately 50 μg of exosomal protein, as determined using a bicinchoninic acid (BCA) assay kit, was purified from differentiated C2C12 cells grown in 100 mL of serum-free cultured Dulbecco’s minimal essential medium on 10 100-mm dishes for 24 hours. In particular, since culture media contain phenol red and amino acids such as tryptophan, tyrosine, and cysteine, which interfere with protein estimation using the BCA assay, culture media must be completely removed from the exosomal pellet in order to accurately determine the protein amount. In addition, serum-free media or exosome-free serum is required for exosomal preparation from cell culture media. Differential centrifugation is a relatively simple method, whereas determination of the morphology and purity of prepared exosomes requires transmission electron microscopy (TEM), the immunoblotting of exosomal markers (e.g., Alix, TSG101, and tetraspanins), and measurement of the size distribution of prepared particles. To rule out contamination by nonexosomal particles, the absence of nonexosomal markers (e.g., GM-130) should be investigated. Further, differential centrifugation protocols according to the matrix of exosome origin should be standardized to minimize variations in exosome purity and content.
Density-Gradient Ultracentrifugation
A possible drawback of differential centrifugation is co-sedimentation of protein aggregates or copurification of proteins that bind nonspecifically to the surface of exosomes. Since exosomes have a buoyant floatation density of 1.08–1.22 g/mL on sucrose gradients [86], density-gradient ultracentrifugation may be a solution for the separation of large protein aggregates from exosomes. In detail, this method separates exosomes from particles of different densities using a linear sucrose or iodixanol cushion. Sucrose density-gradient centrifugation does not allow for the separation of exosomes from viruses or large microvesicles with a comparable sedimentation velocity, and hence an iodixanol gradient (5%–40%) can be substituted for sucrose [87]. In addition, since low-density vesicles take much longer than 16 hours to reach equilibrium density, a 5%–40% iodixanol gradient can be used instead of sucrose to obtain particles with lower densities. Samples overlaid onto the top of the gradient are then centrifuged at 100,000×g for 18 hours at 4˚C. Specific fractions with different densities become relatively enriched in exosomes, as confirmed by exosomal markers and TEM morphological analysis. This method is the most effective for obtaining high-purity exosomes from biological fluids or cell culture medium, although it is laborious, because it involves multiple steps, and there are multiple plausible sources of preanalytical variability, and it may have limited usefulness in clinical settings.
Size-Exclusion Chromatography
The concept behind size-exclusion chromatography (SEC) is that sample particles move through the SEC column (e.g., qEV column, Izon Science Ltd., Cambridge, MA, USA) at different rates depending on their size. That is, particles larger than the desired size will elute more rapidly, while the target population elutes later. This method allows the passage of intact vesicles with a more regular shape than the vesicles obtained using ultracentrifugation. However, the sample volume should not exceed 10% of the resin volume, making the SEC method less useful for large-volume samples (e.g., urine or cell culture media) due to cost. In contrast, for clinical samples with a limited volume, SEC provides exosomes with high purity based on its selective inclusion of the eluted fractions. Although the relative recovery rate is low, SEC may be appropriate to measure levels of targeted biomarkers in limited-volume clinical samples [88].
Commercial Kits Using Polymer-Based Precipitation
Commercially available kits based on the polymer-based precipitation of exosomes are easy to use, fast, and high-throughput, and enable the isolation of exosomes from low-volume samples (as little as 100 μL of serum). The most commonly used kit is the ExoQuick (System Biosciences, Palo Alto, CA, USA). However, it should be noted that this method provides relatively low purity, with coisolate contaminants such as lipoproteins, albumin, and protein aggregates [88]. The precipitation method can be combined with SEC. Specifically, the Exospin purification kit (Cell Guidance Systems, St. Louis, MO, USA) uses SEC combined with precipitation before SEC-based purification on demand. Recently, clinical studies evaluating the performance of AD biomarkers have applied an ExoQuick kit to isolate blood exosomes and observed excellent diagnostic performance [66,76]. However, purity of the prepared exosomes may be an issue [88,89].
Immunoaffinity-Based Capture
Immunoaffinity-based capture is a relatively simple method based on the highly selective separation of exosomes using specific antibodies, such as antiepithelial cell adhesion molecule antibody. Using immunoaffinity-based Dynabeads magnetic separation technology, exosomes can be selectively isolated. However, the selectivity of an antibody targeting a specific subpopulation of exosomes may lead to a lower yield compared to other methods [89]. In addition, the purity of the isolated exosome fraction may be an issue [88,90]. The matrix effects of body fluids will also influence the reactivity and selectivity of the capturing antibody.
CHALLENGES IN THE DEVELOPMENT OF EXOSOMAL BIOMARKERS
Increasing evidence suggests that measurements of the molecular contents of exosomes obtained from body fluids can be used to identify novel non-invasive diagnostic and/or prognostic biomarkers for specific diseases [91]. However, several issues must be resolved to ensure the clinical feasibility of the candidate exosomal biomarkers.
Limited consensus exists regarding the standardization of preanalytical sources of variability, such as patient demographics, body fluid collection, sample handling, sample storage conditions, and conditions/methodology for the preparation of exosomes present in body fluids. Standardization of preanalytical variables is important in exosomal biomarker development, since this is the first step in minimizing variability within and between laboratories. Standardization of the preanalytical process from sample collection to sample storage should be determined according to the type of body fluid, as the unique characteristics of body fluids (e.g., viscosity) or the sample preparation protocols (e.g., centrifugation or use of anticoagulant) may affect the yield and/or size distribution of EVs [92]. In addition, the effects of demographics, freeze-thaw cycles, and storage temperature and duration on the characteristics of exosomes should be determined [90]. The effects of a single freeze-thaw cycle (up to a 1-hour time delay between blood collection and plasma preparation, followed by storage of plasma for up to 1 year at -80°C), were found to be insignificant, whereas freeze-thaw cycle and storage duration were sources of variability in the concentration of EVs in urine and saliva [93]. CSF is an important body fluid that reflects pathologies in the brain. However, there is limited consensus regarding the preanalytical standardization of CSF collection, preparation, and storage. Effects of short-term storage at room temperature and the freeze-thaw cycle on the amount of EVs and abundance of miRNAs in EVs isolated from the CSF of glioblastoma patients have been reported [94]. Therefore, a variety of pre-analytical sources of variability in individual body fluids should be determined, characterized, and standardized to achieve consistent measurement of biomarkers in exosomes.
As described above, there are several methodologies capable of isolating exosomes from body fluids. Each methodology has both advantages and disadvantages in terms of preparation (Table 2), and thus no reference method for exosome purification has been established. This lack of consensus not only affects the isolation of exosomes but also microvesicle preparation from different types of body fluids. Further, size and density overlap between exosomes and microvesicles, making it hard to clearly distinguish between these types of EVs by ultracentrifugation or density-gradient separation. However, density-gradient centrifugation has been used to isolate pure exosomes from samples in vitro (e.g., concentrated cell culture medium). In addition, the biochemical and physical properties of body fluids are diverse, and contamination with non-vesicular materials or structural changes promote the aggregation of exosomes during preparation. None of these issues is specific to exosomes, although the aforementioned exosomal markers have been frequently reported. To estimate the purity of exosomes, both qualitative and quantitative approaches, including morphological observation using TEM analysis, western blotting, or size distribution using dynamic light scanning analysis, can be applied. A consensus paper that proposed a ‘minimal experimental requirement’ for defining EVs was published [95] based on discussion by the Executive Committee of the International Society for EVs. They proposed several criteria that represent the minimal characterization of EVs that should be reported by investigators. For exosomal RNA research, a position paper was also published [96]. The minimal requirements to claim the presence of EVs in isolates are as follows: (1) EVs are isolated from extracellular fluids, that is, from conditioned cell culture medium or body fluids. (2) Several ‘exosome-enriched’ proteins (3 or more) are present in the EV isolates as determined at least semiquantitatively. In addition, the absence of proteins not expected to be enriched in EVs (e.g., intracellular proteins) should be verified. (3) Characterization of single vesicles using at least 2 different technologies (e.g., TEM analysis for morphology and nanoparticle-tracking analysis for size distribution) should be performed. More importantly, detailed experimental protocols from sample preparation to EV isolation as well as their characteristics should be presented in a publication so that other investigators can reproduce the results.
CONCLUSIONS
Recent advances in the discovery of disease biomarkers in EVs, particularly exosomes, support the hypothesis that both EVs and their molecular contents are active players in the pathogenesis of various diseases. Moreover, several biomarkers in EVs have shown reliable clinical performance as diagnostic or prognostic biomarkers in relatively small-scale clinical cohorts of various major diseases (see above). In addition to the development of drug delivery systems using EVs, several ongoing large-scale clinical studies for biomarker development using exosomes in body fluids are underway (www.clinicaltrials.gov). Circulating exosomes are good sources of disease biomarkers, although these are combined exosomes from whole cells and are thus unlikely to be specific to certain cell types. For example, circulating exosomes released from neural cells such as neurons, astrocytes, or microglia are mixed with exosomes originating from other cells in plasma. Therefore, extracting cell-specific exosomes from blood with high purity and in large quantity might improve the specificity of exosomal biomarkers for disease diagnosis. Urinary EVs are the most useful source reflecting molecular events in the genitourinary system, so urinary EVs may be a better source of biomarkers than circulating EVs for urological diseases, including renal diseases, prostate cancer, and bladder cancer. Although the molecules in EVs are diverse and have potential as disease-specific biomarkers, several unresolved issues such as the isolation of pure exosomes from body fluids containing heterogeneous types of EVs should be addressed. Quantification of biomarkers cannot be conducted in clinical settings unless the method of biomarker quantification is repeatable, precise, and accurate. Therefore, to develop a standardized method for isolating EVs, standardized protocols that minimize sources of preanalytical variability, including preparation and storage conditions of body fluids, should be further developed. Such tools will be attainable with additional knowledge of the physiology and components of EVs in diverse contexts, which will accelerate the identification of valuable biomarkers.
Notes
Fund/Grant Support
This work was financially supported by the Mid-Career Researcher Program (2016R1A2B4008399) through the National Research Foundation of Korea (NRF) funded by the Ministry of Science, ICT and Future Planning (J-H Kang, S Kim).
Conflict of Interest
No potential conflict of interest relevant to this article was reported.